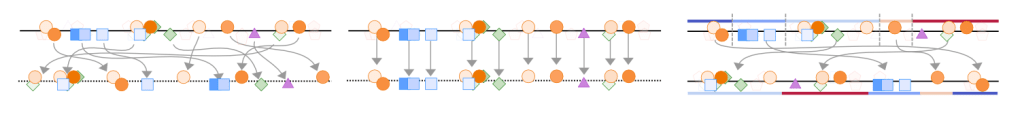
Overview
Biology in multi-cellular systems happens dynamically in space-time. These complex processes are driven in large part as a result of differential expression of genes. Across tissues and cell types, protein production varies dramatically even though each cell of our body harbors the same genomic blueprint. Muscle cells produce motor proteins to effect motion, neurons make ion channels for synapse communication, etc.
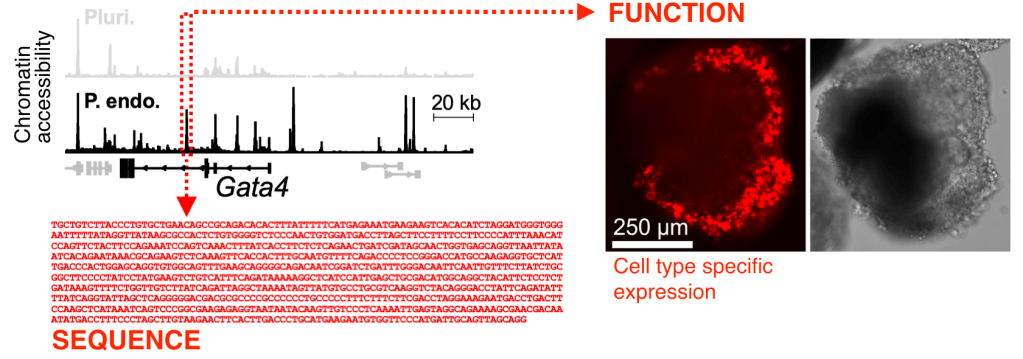
Molecularly, this exquisitely precise control on protein expression is dictated by genetic switches: fragments of non-coding DNA that recruit regulatory proteins which precisely turn genes on and off in development and disease. See an example below of a new endoderm-specific switch we discovered near the gene Gata4.
Our lab addresses a key question of the post-genomic era: how does the DNA sequence of these genetic switches quantitatively encode regulatory output (where, when, how much)? While biophysical and biochemical principles underlying protein and enzymatic functions are relatively well-understood, similar quantitative rules of how these switches functions, called cis-regulatory elements (CREs, also known as transcriptional ‘enhancers’), are still fragmentary.
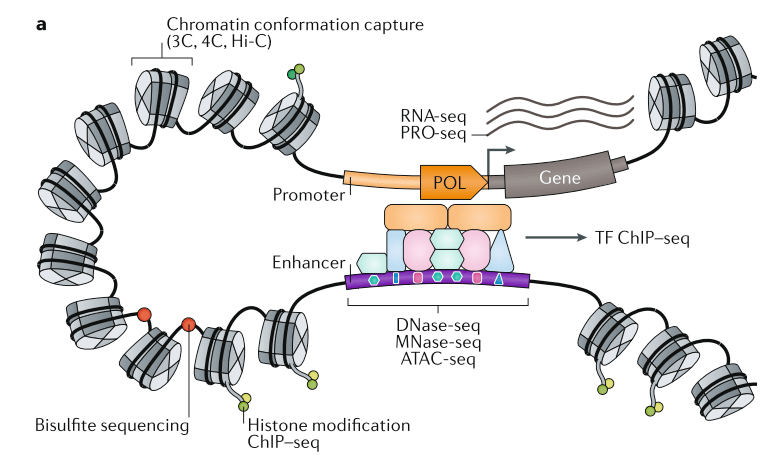
The textbook view, exemplified by the cartoon below (taken from this great review from Gasperini, Tome and Shendure), posits that distal CREs recruit transcription factors and loop to promoters to activate gene expression. But nearly every aspect of this schematic lacks actionable principles. How does the genetics of these regulatory elements work?
We focus on developing new experimental and computational tools to dissect the sequence-to-function maps of these important non-coding elements in various in vitro mammalian models of development and disease.
Key techniques & questions
Single-cell massively parallel reporter assays
We were the first to combine single-cell RNA-seq and barcoded reporters to profile quantitatively the activity of distal CREs in complex multicellular systems. These ‘single-cell quantititative expression reporters’ (scQers) are currently being used to study gene regulation in a variety of in vitro and in vivo model systems. See the paper here, with a concise briefing explaining why we think this technology will be transformative for regulatory biology.
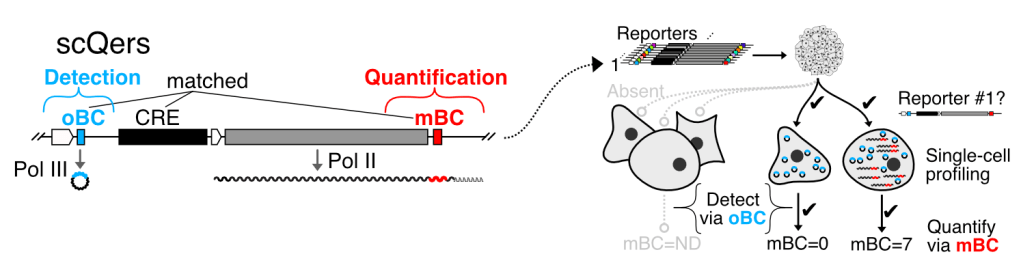
We have used scQers to profile quantitatively in a parallelized manner the activity of cell-type specific enhancers within stem-cell models of early development:
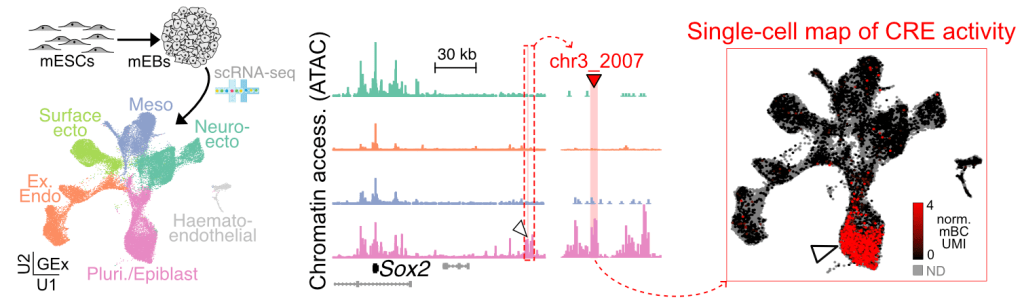
Currently, we develop new applications of scQer in models of cancer progression to precisely engineer regulatory elements targeting specific cell-states with exquisite precision.
Barcoded enhancer assays (MPRA)
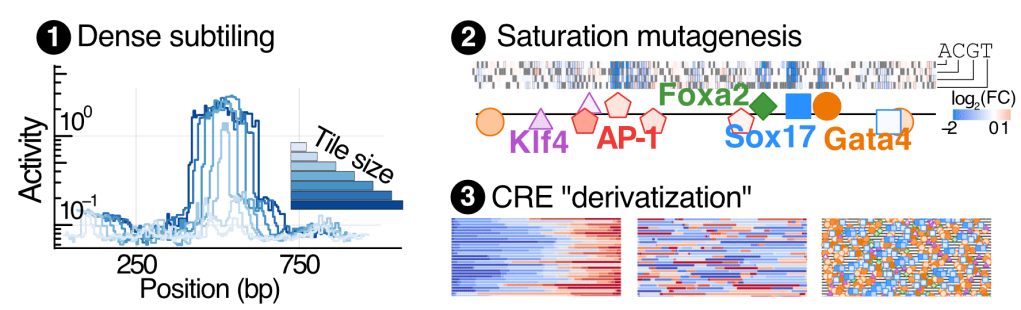
We also use bulk assays to probe various aspects of regulatory sequence-to-function maps quantitatively at ultra-high scale. We focus on under-explored of the CRE design space:
1. Multi-sized CRE sub-tiling to chart the relationship linking CRE size and activity.
2. Saturation mutagenesis to map TF binding sites a single-base-pair resolution.
3. Statistical CRE derivatization to assess flexibility of the regulatory grammar.
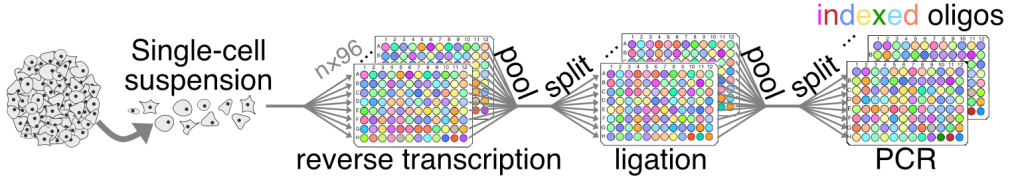
Scalable single-cell RNA-seq
We use an ‘open-source’ optimized, plate-based combinatorial indexing approach, as detailed here, to perform single-cell-seq at a fraction of the cost of commercial platform. The lab has experience leveraging the assay for functional measurements in addition to the usual molecular ‘atlasing’ of multi-cellular samples
Mathematical modeling & model-driven CRE design
We use biophysical and deep-learning based approaches to predict activity from sequences. Our bioinformatic projects span the purely theoretical (e.g., trying to understand biophysical limits of regulatory specificity) to applied (optimizing CRE activity) axes.