Mammalian regulatory genomics (2020-present)
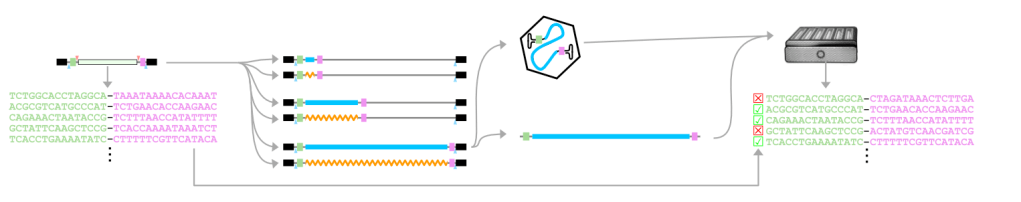
Extensive length and homology dependent chimerism in pool-packaged AAV libraries
J.-B. Lalanne et al, bioRxiv 2025
Multiplex profiling of developmental cis-regulatory elements with quantitative single-cell expression reporters
J.-B. Lalanne*, S. Regalado* et al, Nature Methods 2024
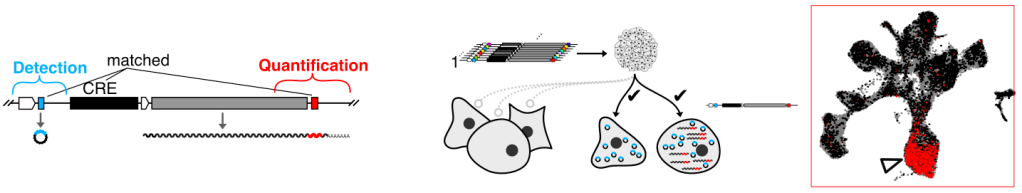
See the associate Research Briefing:
Quantitative profiling of regulatory DNA activity at single-cell resolution
Contributing author
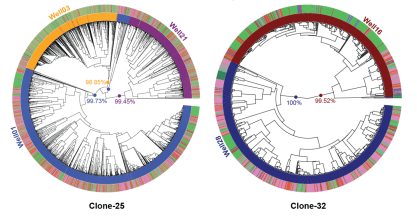
Lineage recording in monoclonal gastruloids reveals heritable modes of early development
S. Regalado* & CX Qiu* et al, bioRxiv 2025
Barcoded monoclonal embryoids are a potential solution to confounding bottlenecks in mosaic organoid screens
S. Regalado* & CX Qiu* et al, bioRxiv 2025
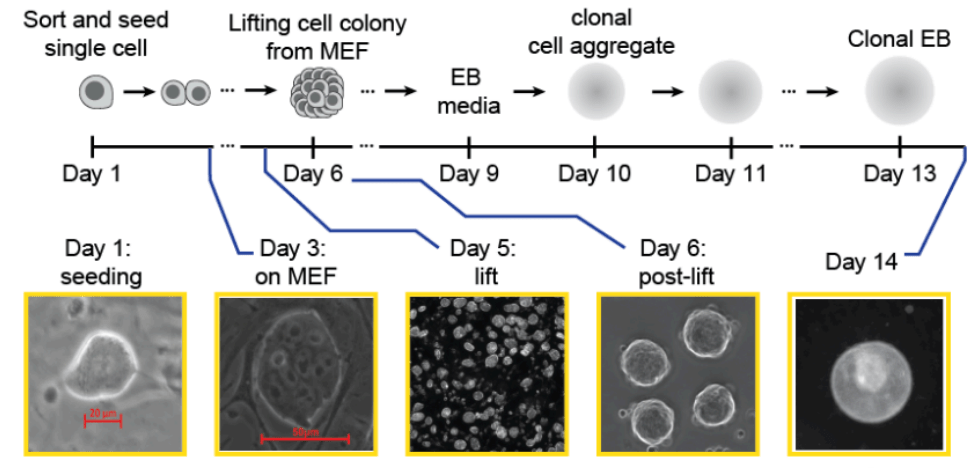
Multiplex generation and single-cell analysis of structural variants in mammalian genomes
S. Pinglay et al, Science 2025
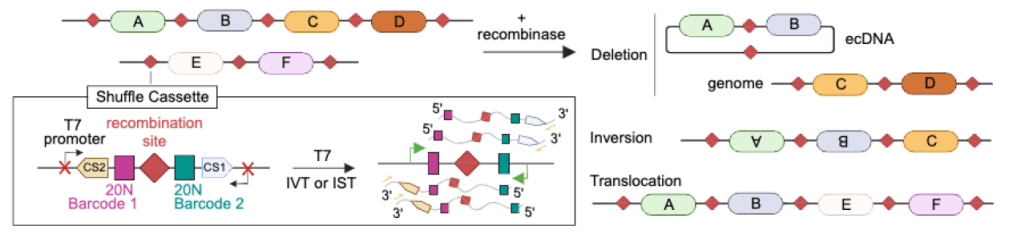
Technical and biological sources of noise confound multiplexed enhancer AAV screening
A. Hunker et al, bioRxiv 2025
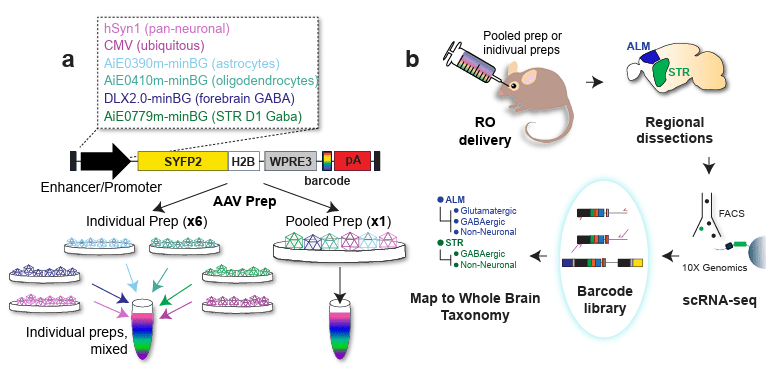
Multiplex, single-cell CRISPRa screening for cell type specific regulatory elements
F. Chardon* & T. McDiarmid* et al, Nature Communication 2024
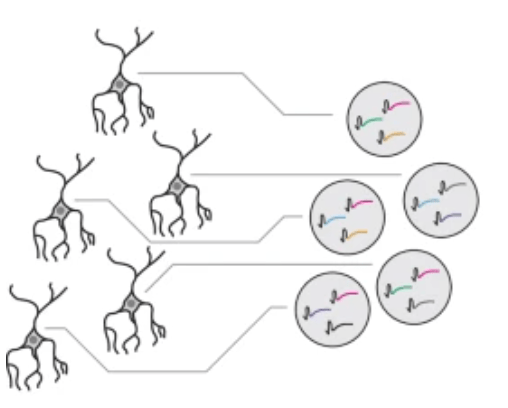
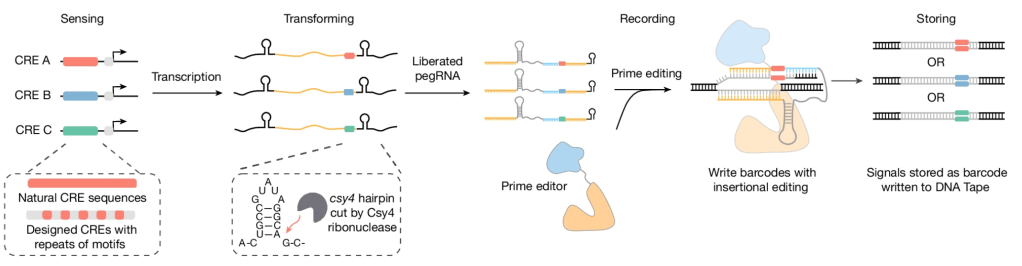
Symbolic recording of signalling and cis-regulatory element activity to DNA
W. Chen et al, Nature 2024
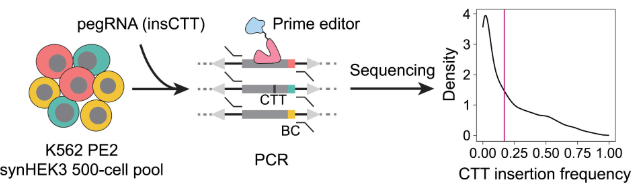
Chromatin context-dependent regulation and epigenetic manipulation of prime editing
X. Li et al, Cell 2024
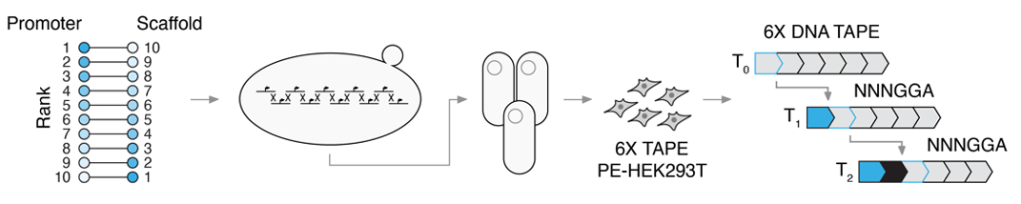
Diversified, miniaturized and ancestral parts for mammalian genome engineering and molecular recording
T. McDiarmid* & M. Taylor* et al, bioRxiv 2024
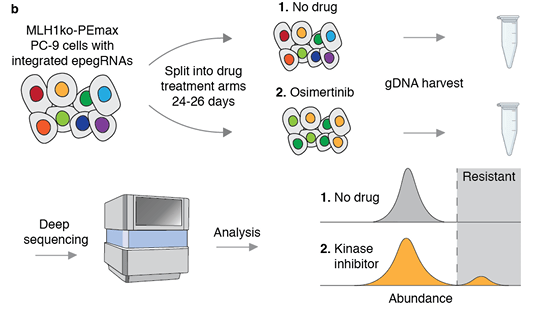
A multiplex, prime editing framework for identifying drug resistance variants at scale
F. Chardon et al, bioRxiv 2023
Quantitative bacterial gene regulation (2015-2020)
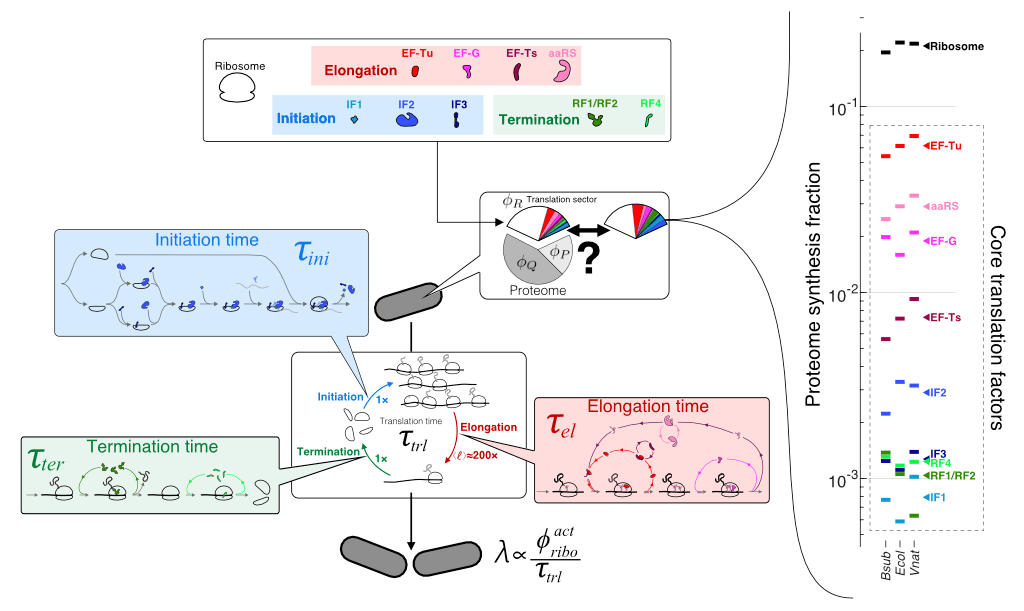
First-principles model of optimal translation factors stoichiometry
J.-B. Lalanne & G.-W. Li, eLife 2021.
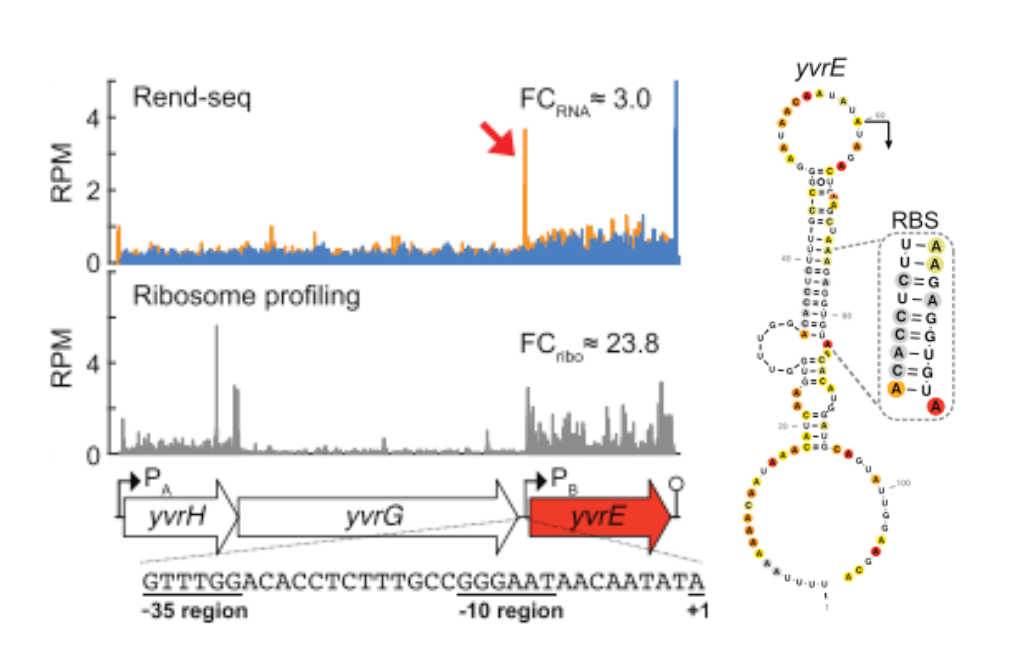
Differential translation of mRNA isoforms transcribed with distinct sigma factors
D. McCormick* & J.-B. Lalanne* et al, RNA 2021.
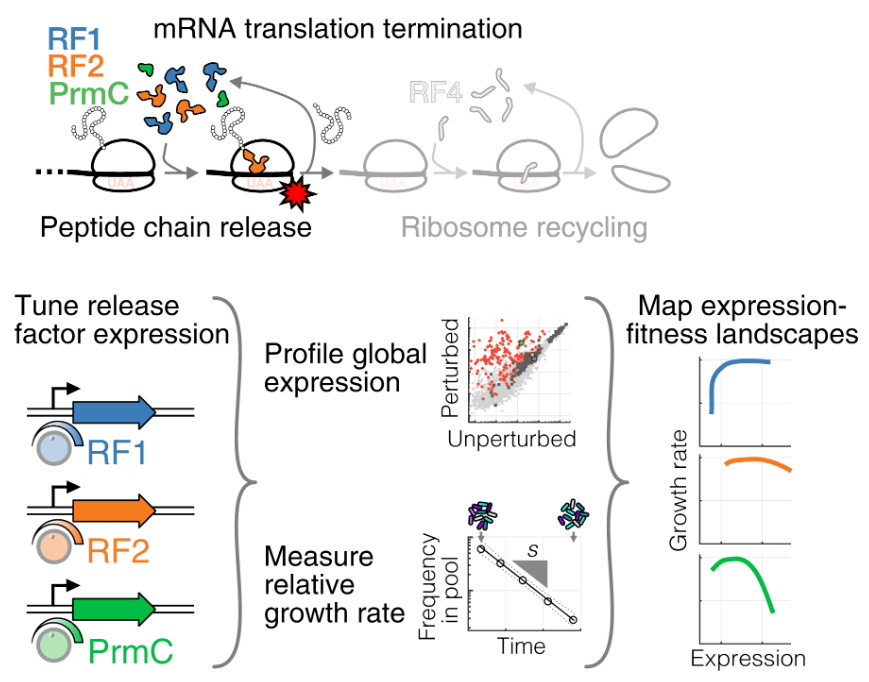
Spurious regulatory connections dictate the expression‐fitness landscape of translation factors
J.-B. Lalanne, D. Parker, and G.-W. Li, Molecular Systems Biology 2021.
Quantitative Control for Stoichiometric Protein Synthesis
J. Taggart*, J.-B. Lalanne*, and G.-W. Li, Annual Review of Microbiology 2021.
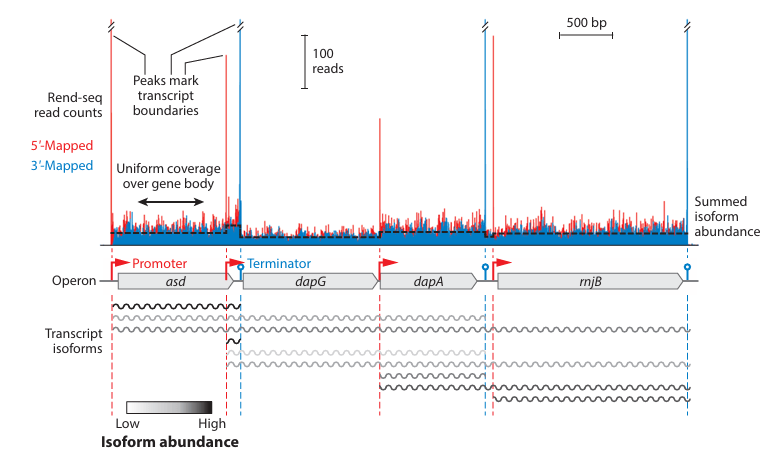
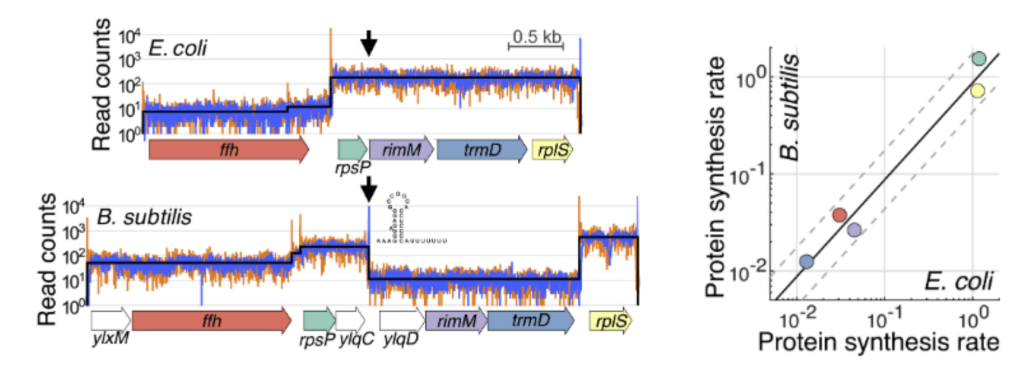
Functionally uncoupled transcription–translation in Bacillus subtilis
G. Johnson*, J.-B. Lalanne*, M. Peters, and G.-W. Li, Nature 2020.
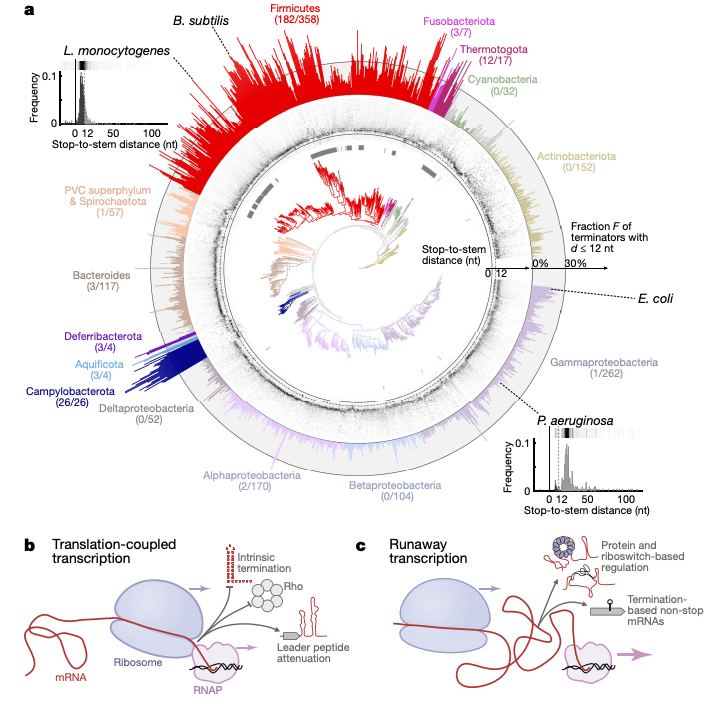
Evolutionary convergence of pathway-specific enzyme expression stoichiometry
J.-B. Lalanne et al, Cell 2018.
Contributing author
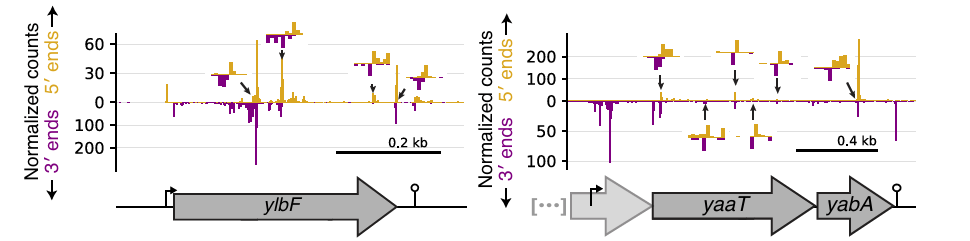
A high-resolution view of RNA endonuclease cleavage in Bacillus subtilis
J. Taggart et al, NAR 2025.
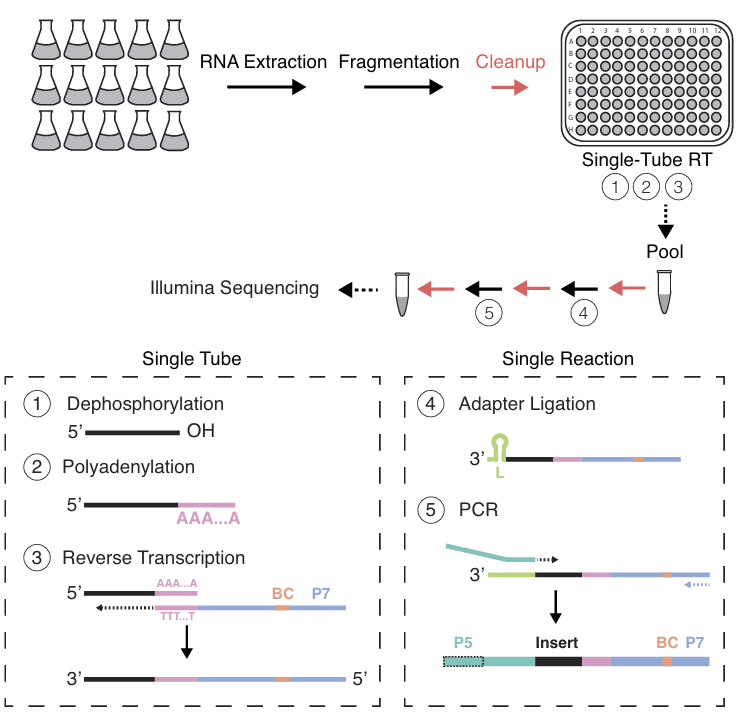
BaM-seq and TBaM-seq, highly multiplexed and targeted RNA-seq protocols for rapid, low-cost library generation from bacterial samples
G. Johnson* & D. Parker* et al, NAR 2023.
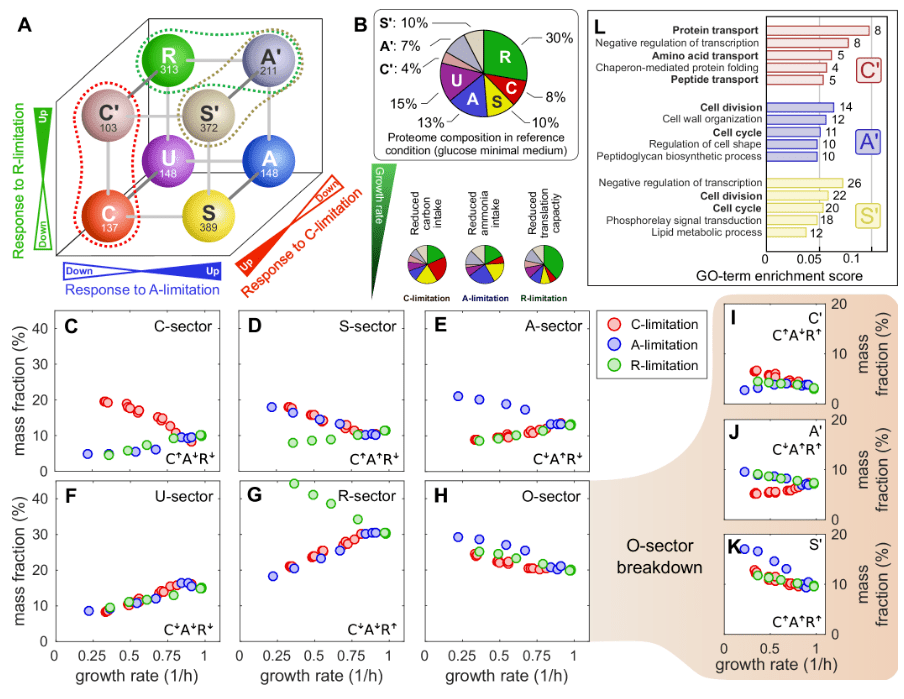
From coarse to fine: the absolute Escherichia coli proteome under diverse growth conditions
M. Mori et al, Molecular Systems Biology 2021.
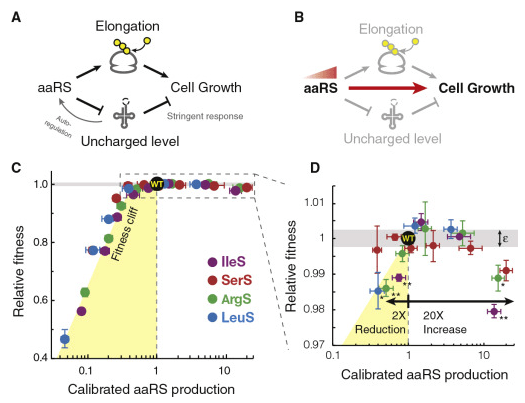
Growth-optimized aminoacyl-tRNA synthetase levels prevent maximal tRNA charging
D. Parker et al, Cell Systems 2020.
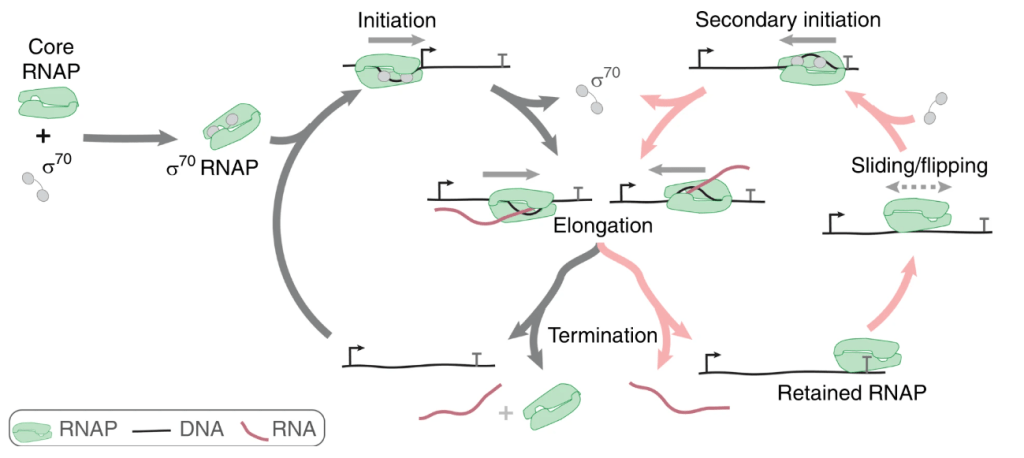
Alternative transcription cycle for bacterial RNA polymerase
T. Harden et al, Nature Communications 2020.
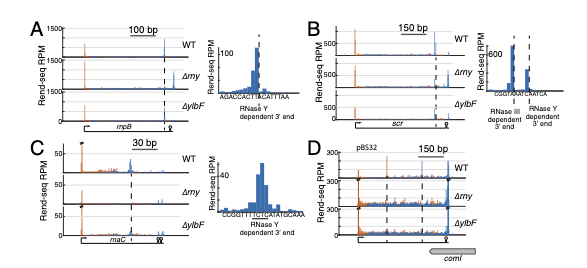
Maturation of polycistronic mRNAs by the endoribonuclease RNase Y and its associated Y-complex in Bacillus subtilis
A. DeLoughery et al, PNAS 2018.
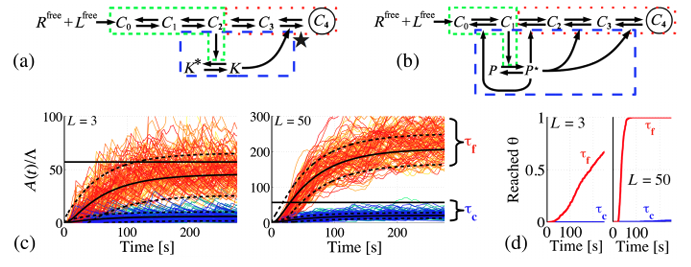
Theoretical biophysics of chemo-sensing (2012-2014)
Chemodetection in fluctuating environments: Receptor coupling, buffering, and antagonism
J.-B. Lalanne and P. François, PNAS 2015.
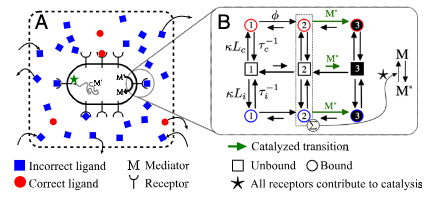
Principles of Adaptive Sorting Revealed by In Silico Evolution
J.-B. Lalanne and P. François, Physical Review Letters 2013.